Antibody validation Practical guide to finding and validating suitable antibodies for research
Example step 1 How to define a chosen antigen for a monoclonal antibody to PIM2
1 Identify the target antigen
It is essential to identify the ‘approved nomenclature’![]() For any protein comprises a short-form abbreviation known as its gene symbol and a longer more descriptive name. The HUGO Gene Nomenclature Committee (HGNC) ensures that each gene is given only one approved gene symbol. This unique identifier allows the clear and unambiguous referencing of genes in scientific communications, and facilitates any electronic data retrieval from databases and publications of the target protein to obtain correct information about the protein and any specific antibody or antibodies that may be available. There are several different research engines for identifying the 'approved nomenclature', two of which are listed below.
For any protein comprises a short-form abbreviation known as its gene symbol and a longer more descriptive name. The HUGO Gene Nomenclature Committee (HGNC) ensures that each gene is given only one approved gene symbol. This unique identifier allows the clear and unambiguous referencing of genes in scientific communications, and facilitates any electronic data retrieval from databases and publications of the target protein to obtain correct information about the protein and any specific antibody or antibodies that may be available. There are several different research engines for identifying the 'approved nomenclature', two of which are listed below.
Working example: PIM2
- a) Using GENECARDS website the alternative names/aliases of PIM2 are: pim-2 oncogene; Pim-2h; PIM2; proto-oncogene Pim-2 (serine threonine kinase); serine/threonine protein kinase pim-2; Serine/threonine-protein kinase pim-2.
- b) Using GENENAMES the alternative names/aliases of PIM2 are: Pim-2 proto-oncogene and serine/threonine kinase.
2. Confirming the protein sequence
Identifying the "canonical" protein sequence is another essential step permitting the collection of accurate information about the target protein. Examples of such information include its molecular weight and the presence of any known variants, e.g. those produced as a result of alternative splicing, proteolytic cleavage, post-translational modification. Use the Uniprot website to identify PIM2 protein sequence.
Working example: PIM2
A video demonstrating how the PIM2 canonical sequence can be obtained from the Uniprot webpage is shown below:
Result from the Uniprot search:
MLTKPLQGPPAPPGTPTPPPGGKDREAFEAEYRLGPLLGKGGFGTVFAGHRLTDRLQVAI KVIPRNRVLGWSPLSDSVTCPLEVALLWKVGAGGGHPGVIRLLDWFETQEGFMLVLERPL PAQDLFDYITEKGPLGEGPSRCFFGQVVAAIQHCHSRGVVHRDIKDENILIDLRRGCAKL IDFGSGALLHDEPYTDFDGTRVYSPPEWISRHQYHALPATVWSLGILLYDMVCGDIPFER DQEILEAELHFPAHVSPDCCALIRRCLAPKPSSRPSLEEILLDPWMQTPAEDVPLNPSKG GPAPLAWSLLP
Figure legend Uniprot PIM2 canonical sequence showing the aminoacid sequence.
3. Identify the existence of variants/isoforms
The isoforms and variants of the protein under investigation can be identified from the following websites:
The information obtained from these websites enables the user to determine the existence of possible variants and isoforms of the target protein and to decide which of these need to be detected in the relevant experimental system(s). For instance, multiple variants of a protein may result in the presence of additional bands in Western blots or altered patterns of tissue distribution and/or subcellular localisation in immunohistochemical studies that might otherwise be mistaken for non-specific reactivity.
Working example: PIM2
The ACEVIEW website predicted at least six spliced PIM2 variants and seven different isoforms (4 complete, 2 COOH complete and I partial) as well as functional domains. An example of the results obtained from this website are shown below.

Figure legend The mRNA map from ACEVIEW predicting the presence of at least six different variants of PIM2.
A PubMed search showed the existence of 3 different PIM2 isoforms of 34 kDa, 37 kDa and 40 kDa. This information, besides supporting the presence of more than one PIM2 protein also indicates that any antibody directed against a region of PIM2 common to all three isoforms would produce a Western blot showing three different bands rather than one.
4. Investigate the presence of related proteins to the target antigen
Sequence analysis of the target antigen should be performed, using the NCB1 protein BLAST tool, that will identify other proteins (in the species under study) that share significant sequence identity with the whole or part of the target protein.
Working example: PIM2
PIM proteins are a family of short-lived serine/threonine kinases that are highly conserved through evolution in multicellular organisms. PIM-2 is highly homologous to PIM1 and PIM3 proteins and all three PIM proteins possess similar oncogenic functions. This means that PIM2 reagent(s) raised against a full-length protein or an unknown PIM antigen may also cross-react with PIM1 and PIM3 proteins. It is, therefore ESSENTIAL to check any cross reactivity by performing a sequence search using the protein reference sequence. The video shows how this can be done for the PIM2 protein.
Result from the BLAST search:
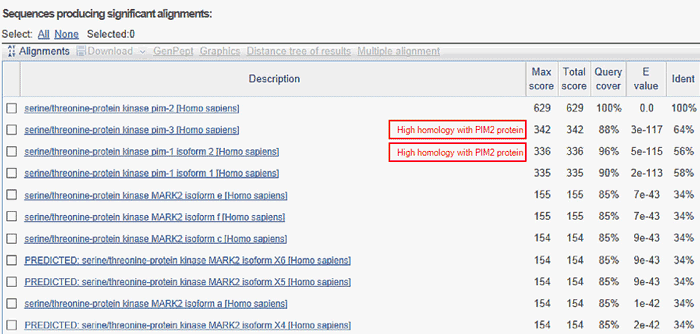
Figure legend Results from a BLAST search of PIM2 showing related proteins and the degree of similarities between their aminoacid sequences. PIM3 and PIM1 share 64% and 56%, respectively, identity with PIM2 (boxed). This means that, if an antibody has been raised using a full-length PIM2 protein or a part of the protein that is highly homologous to all three PIM family members, there is a very high probability that the antibody also detects PIM1 and PIM3 proteins in addition to PIM2.
5. Define ideals epitopes
A BLAST search can also assist in the identification of ideal epitopes that could be used for antibody production. These aminoacid regions should be unique to the target protein. The Clustal Omega Multiple Sequence Alignment programme can then be used to identify alignments specific for the target molecule.
Working example: PIM2
The video below demonstrates the use of Clustal Omega Multiple Sequence Alignment programme for PIM2.
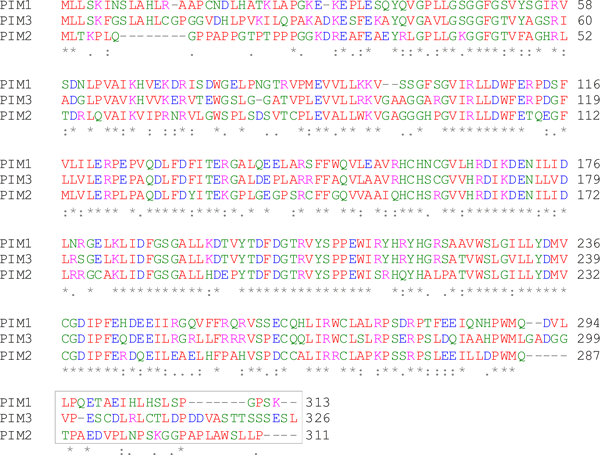
Figure legend Alignment of the three PIM family members to show regions of PIM2 to produce an immunogen suitable for raising an antibody specific for PIM2. Aligning the protein sequences of all three PIM family proteins shows that the C-terminal portion (boxed) of the PIM2 protein is the best region for producing an antibody that should be specific for the PIM2 protein only.
